Programma Erasmus+ "LIVE ON THE KARST"
Attività progetto a.s.2016-2017
Our invisible friends
Nell’ambito della disciplina “Microbiologia e Biotecnologie Ambientali” la Prof.ssa Antonella Serafin ha coordinato l’attività di ricerca della classe IV ABA, lavorando a partire da delle patine microbiche isolate nella grotta di Planina in Slovenia, durante la prima sessione internazionale di apprendimento (Aprile 2016).
Summary/Abstract
During our 1st learning session (11-15 April 2016) in Sezana, Slovenia, we visited Planina Cave (Planinska jama) on Wednesday, 13th April.
Even if the purpose of the visit was to admire the largest water-filled cave in Slovenia, on the Southern edge of the Planina plain, our attention was captured by the golden effect on some of the walls known to be formed by some microbial organisms.
With the kind help of our guide two samples have been taken using sterile swabs from some/a reachable (in terms of height) wall(s). The site was attended by bats.
Four strains have been isolated in the “Levi-Ponti” Microbiology lab. All of them have been sent to DAFNAE, the University of Padova’s Department of Agronomy, Food, Natural Resources, Animals and the Environment, where 16 S region has been extracted and amplified. Only two of them were successfully processed. The two amplified region were sent to an external contract lab for sequencing.
Introduction
It is one of the largest Slovenian caves including the interesting feature of the subterranean confluence of the Pivka and Rak rivers. Beneath a picturesque 100 meter high cliff lies one of the mightiest cave springs in Slovenia. Planina cave is also known for its fauna: bats, Proteus anguinus and the tiny, blind, transparent crabs.
Some of the cave walls’ sides look iridescent if lit with a torch, similar to pyrite minerals. This effect is given by very small water drops condensed on a soft substrate of soil granules where lots of microbial organisms can settle. They are described as “cave gold” (1). The golden aspect of organic layer and microbial colonies usually appears when illuminated water droplets magnify the yellowish pigment of the microbial mat beneath the water film (Mulec, 2008).
One of “Live in Karst” Erasmus+ project objectives is presentation of research results from environmental surveys and sharing of teaching methods during learning.
Materials and Methods
Isolation
Two surface swab samples (10 cm2) were taken aseptically in caves from “golden aspect” wall surface. The removed coat appeared to be soft, with hyphae and very subtle particles like clay.
The swabs were individually collected and transferred into tubes containing Amies transport medium without charcoal (Kima 230397, Piove di Sacco, Padova, Italy - Fig. 1) and delivered at room temperature to Solski centre Srecka Kosovela biology laboratory (Sezana) within 5 hours. There they have been stored between 2-8 °C for 48 hours and then transported to Italy at room temperature within 5 hours, where they were finally placed into refrigerated conditions (for 60 hours) within the laboratory for testing.
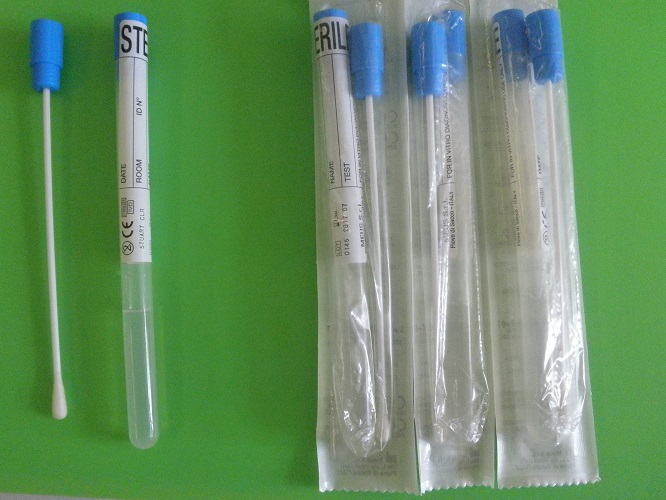
Fig. 1: Swabs and their tubes containing Amies transport medium without charcoal (Kima 230397, Piove di Sacco, Padova, Italy), opened and entirely sealed.
On Monday 18th April each sample was aseptically suspended into a 10 ml tube of Nutrient Broth.
Preparation of dilutions: 9 ml volumes of Nutrient Broth were dispensed in four loose capped tubes. The undiluted swab sample was mix by vortex. With a straight-sided pipette dipped in only half an inch, 1 ml was removed. It was delivered into the first dilution blank, about half an inch above the level of the liquid. The pipette was discarded. The tube was capped and then mixed by vortex to avoid aerosol formation. 1 ml was removed with a fresh sterile pipette and transferred to the next dilution blank. Operations were repeated for the required number of dilutions according to the following scheme:
Tube no. 1 2 3
Dilution 1 : 10 1 : 100 1 : 1000
Vol of original fluid/ml 0.1 0.01 0.001
(or 10-1) (or 10-2) (or 10-3)
Growth conditions
The broths were pre-incubated at room temperature for a couple of hours, it not being possible to keep them at the average temperature (8-11 °C) of the Planina cave (Fig. 2).

Fig. 2: Some of the tubes after pre-incubation and inoculation.
Following pre-incubation, 1-ml of each undiluted and diluted culture was pipetted into the centre of the appropriate sterile Petri dish using a gamma-irradiated sterile plastic pipette.
15 ml of melted Mueller – Hinton agar were added and mixed to each dilution according to the plate count technique. Operations were repeated for the same cultured samples using Mueller – Hinton agar supplemented with oxytetracycline, an antibiotic that increases the counts of yeasts and moulds. A 10-ml suspension of lyophylised antibiotic was added into 500 ml of melted M.H. agar.
Both the two sets of plates (with and without antibiotic) were incubated at room temperature for 48-72 h.
RESULTS
After 48-72 hours all the M. H. agar plates showed the growth of a very high number of microorganisms, different in variety, while on the plates supplemented with oxytetracycline only few colonies were able to grow according to Table 1 and Fig. 3
|
Plate count from swab into N. B. broth |
||
|
Dilution |
M.H. Agar |
M.H.A.+ ANTIBIOTIC |
|
1 : 1 |
positive growth N.A. |
18 CFU/ml |
|
1 : 10 |
positive growth N.A. |
80 CFU/ml |
|
1 : 100 |
positive growth >>300 |
- |
|
1 : 1000 |
positive growth N.A. |
- |
Table 1: Results yielded from plate count technique. N.A. stands for Not Applicable, when some colonies are confluent or spread too much on the plate so that they hide other colonies.

Fig. 3: Difference in the growth results at different dilution without and with antibiotic.

Fig. 4: Colonies growth on M.H. agar plates opened inside biohood


Fig. 5: Oxytetracycline resistant microorganisms growth (dilution 1:10)
Several colonies from M.H agar plates have been selected and subcultured on the same agar medium or onto Tryptone Soya Agar and Columbia agar plates in order to isolate pure strains (Fig. 6).


Fig. 6: some example of isolated strains on different culture media agar plate
Finally eight strains have been isolated from M.H.A. agar plates and we made a Gram stain for each of them. The strains were mostly Gram-, short-rod bacilli (Figs. 7 and 8) and one curved bacillus Gram+ strain (Fig. 9)

Fig. 7: Strain 4 B magnified with Optical Microscope 400X (Gram stain and picture taken by M. Canal)

Fig. 8: Gram- bacilli Optical Microscope X 1000 with oil (Gram stain and picture taken by E. Bagaggiolo)

A

B
Fig. 9: Gram+ rods at 1000X magnification (A) and at 400X (B) (Gram stain and picture taken by S. Pavan)
From M.H. agar supplemented with oxytetracycline some moulds have been isolated (Fig. 10) and a very peculiar microorganism (Figs. 11, 12 and 13).

Fig. 10: Mould Gram stain at 1000X magnification (Picture taken by V. Libralesso)
10 days after the swab were examinated by pour plate method on M.H. agar supplemented with oxytetracycline, one of the subcultured colonies formed a white aerial mycelium (Figs. 11 and 12).
The strain was subcultured (Figs. 13, 14 and 15) in order to perform a good Gram stain cell. This operation wasn’t easy because the colony was dry and lots of cells stayed attached to the loop. One of the best Gram stain results is shown below (Figs. 16, 17 and 18).

Fig. 11

Fig. 12

Fig. 13

Fig. 14

Fig. 15: Strain 1PX

Fig. 16: strain 1 PX (100 X) Foto by A. Carraro

Fig. 17: strain 1 PX (400 X) Foto by A. Carraro

Fig. 18: Strain “1PX” (1000 X)
Four strains were preserved by making thick suspensions of bacteria in commercially available vials and then freezing at – 20°C. In this commercial system, suspensions are added to vials containing prepared beads. The excess fluid is removed and the vial frozen. One bead is removed to culture medium as required (Figs. 19 and 20).

Fig. 19

Fig. 20: Strain 2A after one year of -20 °C storage
Bacterial Identification
At the beginning of July, these four strains were sent to DAFNAE, the University of Padova’s Department of Agronomy, Food, Natural Resources, Animals and the Environment which is situated in Legnaro (PD).
This has been possible thanks to Prof. ssa Marina Basaglia and Prof. Sergio Casella and their commitment to education.
At the DAFNAE microbiology laboratory, DNA was extracted from isolated colonies using the standard protocol for genetic bacterial identification and characterization. This purified DNA was used for PCR of 16S rDNA. Fragments were firstly analyzed by 1% agarose gel electrophoresis yielding the results shown below (Fig. 21) demonstrating DNA extraction and 16S amplification. Unfortunately only strains 2A and 4B resulted positive for the presence of amplified DNA from region 16S. The other strains probably needed further methods of DNA extraction.

Fig. 21: Gel electrophoresis for PCR of 16S rDNA (kindly donated by J. E. Rodríguez)
These two strains 16S rDNA sequences were amplified and sent to BMR Genomics (spin off of the University of Padova) for sequencing.
16S rDNA sequences were determined and analyzed in the GenBank using BLAST search (https://blast.ncbi.nlm.nih.gov). Below is one of the two 16S rDNA sequences obtained:
GGTAGTGGGGGNNANGNTTCGAAAGGAACGCTAATACCGCATACGTCCTACGGGAGAAAGCAGGGGACCTTCGGGCCTNNNNTATCAGATGAGCCTAGGTCGGATTAGCTAGTTGGTGAGGTAATGGCTCACCAAGGCGACGATCCGTAACTGGTCTGAGAGGATGATCAGTCACACTGGAACTGAGACACGGTCCAGACTCCTACGGGAGGCAGCAGTGGGGAATATTGGACAATGGGCGAAAGCCTGATCCAGCCATGCCGCGTGTGTGAAGAAGGTCTTCGGATTGTAAAGCACTTTAAGTTGGGAGGAAGGGTTGTAGATTAATACTCTGCAATTTTGACGTTACCGACAGAATAAGCACCGGCTAACTCTGTGCCAGCAGCCGNGGTAATACAGAGGGTGCAAGCGTTAATCGGAATTACTGGGCGTAAAGCGCGCGTAGGTGGTTCGTTAAGTTGGATGTGAAATCCCCGGGCTCAACCTGGGAACTGCATTCAAAACTGTCGAGCTAGAGTATGGTAGAGGGTGGNGGAATTTCCTGTGTAGCGGTGAAATGCGTAGATATAGGAAGGAACACCAGTGGCGAAGGCGACCACCTGGNCTGATACTGACACTGAGGTGCGANNGCGNGGGGAGCAAACAGGATTAGATACCCTGTAGTCCACGCCGNAAACGATGTCAACTAGCCGTNGGGAGCCTTGAGCTCTTAGTGGCGCAGCTAACGCATTAAGTTGACCGCCTGGGAGTACGGCCGCAAGGNTAAAACTCAAATGAATTGACGGGGGCCCNNACNAGCNGNGNNCATGNGNTTAATTCGAAGCAACNNNNAGAACCTTACCAGGCCTTGACATCCAATGAACTTTCCAGANATGGATTGNNCCTTCGNNNATTNNNANNNNTGCTNCATGCTGNCNNCAGNNTNGTNNCNNNAGATGTGGTNANTCCNNACNNNCGCANCC
Both strains have been identified with certainty at the genus level as Pseudomonas spp. (matching 93 % for strain 2A and 96% for strain 4B for species P. granadensis)

Fig 22: Pseudomonas spp. “2A” on TSA plate isolated at the end of April 2016

Fig. 23: Pseudomonas spp. “2A” Gram stain made by A. Michieli in April 2016 (400X)

Fig. 24: Pseudomonas spp. “2A” Gram stain at 1000X magnification made by S. Pavan in May 2017
One year later, from “Levi-Ponti” Microbiology Laboratory’s stock cultures, the two strains “2A” and “2B” have been revitalized on to TSA plates (Figs. 20 and 25). Gram stains have been performed again yielding the results shown in Figs. 24, 26 and 27.

Fig. 25: Pseudomonas spp. “4 B” on 11th May 2017 (picture taken by M. Canal)

Fig. 26: Pseudomonas spp. “4 B” Gram stain at 400X magnification made by M. Canal

Fig. 27: Pseudomonas spp. “4 B” Gram stain at 1000X magnification made by M. Canal
Discussion
Considering this has been a trial experience begun with materials we had in our laboratory, as a first attempt at isolation the results are satisfactory. Eight different pure culture strains were obtained and four of them were stored and preserved as stock culture. Two of them have been identified with certainty at the genus level as Pseudomonas spp.
Certainly due to the simplicity of isolation methods used we selected only the most common species. Pseudomonads usually are present in soil at the concentration 104 UFC/g.
Acknowledgements
Gena Petric, our guide at Planinska jama
Maja Preseren and all the school team of Gimnazija in Ekonomska Sola (Solski center Srecka Kosovela Sezana)
Prof. ssa M. Basaglia and Prof. S. Casella (DAFNAE - Department of Agronomy Food Natural Resources Animals and Environment, University of Padova)
Jesús E. Rodríguez G. (DAFNAE - Department of Agronomy Food Natural Resources Animals and Environment, University of Padova)
Aurora Masiero and Tiziana Stocco (laboratory assistants at I. I. S. “Primo-Levi”)
Danieli, E. Rocco, A. Pastrello and A. Zamengo students from V ABA at I. I. S. “Primo-Levi”, school year 2015-16
References
Collins C. H., Lyne P. M. and Grange J. M., 1995 – Microbiological Methods, VII Edition, BH Ltd.
Mulec J., 2008 - Microorganisms in hypogeon: examples from Slovenian karst caves. Acta Carsologica, 37: 153-160.
Mulec J., Oarga-Mulec A., Tomazin R. and Matos T., 2015 - Characterization and fluorescence of yellow biofilms in karst caves, southwest Slovenia - International Journal of Speleology 44 (2) 107-114, May 2015
Pasić L, Kovce B, Sket B, Herzog-Velikonja B., 2009 - Diversity of microbial communities colonizing the walls of a Karstic cave in Slovenia - FEMS Microbiol Ecol 71 (1): 50-60.